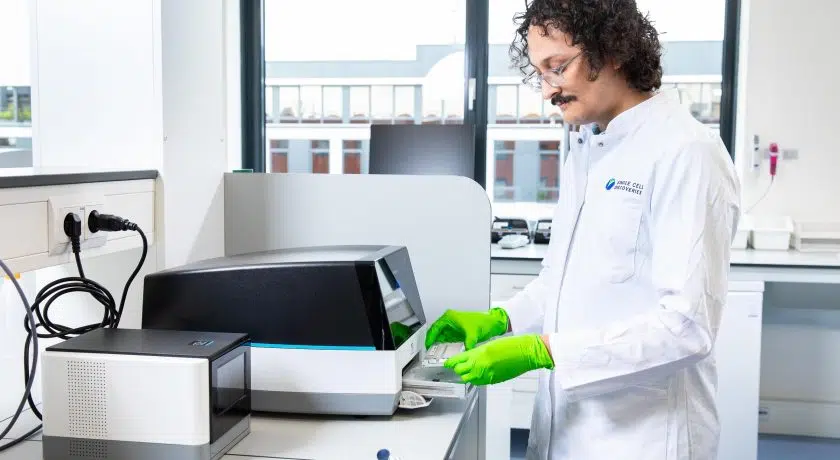
This post describes what single-cell ATAC-seq essentially is and what you can do with it.
Single-cell RNA sequencing is a powerful way to determine a tissue’s properties and dynamics in health and disease. However, it can be valuable to gather more than a cell’s gene expression profile to fully understand the state of cells during illness or treatment.
Indeed, we observe a growing demand among life science researchers for single-cell technologies that unearth additional features. The market offers technologies focusing on such features, like a cell’s set of proteins, metabolic states, or epigenetic profiles.
Chief among the latter type is single-cell ATAC-seq (scATAC-seq). This technology enables researchers to capture the chromatin accessibility profiles of tissues in single-cell resolution. With these profiles, researchers can determine how tissues change during disease and react to compounds based on their chromatin accessibility.
Not unlike single-cell RNA sequencing, scATAC-seq is a scalable, high-resolution assay with many possibilities. Scientists from various fields have thus adopted the technology, from developmental biology and immunology to cancer research and drug discovery. Single Cell Discoveries has also added the technology to its portfolio and is transferring its value to clients around the world. In this blog, we describe the fundamentals of what scATAC-seq is and how you can employ it.
Read on to learn more about:
- The essentials of single-cell ATAC-seq
- The name explained
- What it enables you to do
- scATAC-seq at Single Cell Discoveries
scATAC-seq essentials
At its core, single-cell ATAC-seq captures information on the chromatin accessibility profile of individual cells. It is the single-cell variant of bulk ATAC-seq, now one of the most popular approaches to investigate epigenetic profiles of tissues.
ATAC-seq was developed in 2013 by the Greenleaf group of the Stanford University School of Medicine. Two years later, the group published the single-cell version of the technology. Its protocols and reagents have been made commercially available by several companies, including 10x Genomics, BioRad and Active Motif.
What is chromatin?
In a nutshell, chromatin is DNA wrapped around histone proteins and forms the chromosomes found in our cells. It packages the DNA of our genome into a highly compact form that can fit in the cell nucleus.
When packaged tightly, the chromatin is closed for transcription. When it is open, gene expression is possible. Hence, chromatin accessibility often correlates with active gene expression.
DNA is wrapped around histone proteins to create chromatin. “Open” chromatin is generally open for transcription factor binding, leading to gene expression.
For an up-to-date and comprehensive overview of the effects of chromatin state and other types of genome regulation, see this review in Nature Reviews Genetics.
Chromatin accessibility profile
In biology research, Chromatin accessibility often translates into the availability of a stretch of DNA for gene expression regulators such as transcription factors. Transcription factors mount on available DNA to guide the pileup of the transcription machinery that further enables gene expression. Chromatin accessibility, thus, is tightly connected to gene regulation and expression.
The chromatin accessibility profile shows the genome-wide map of accessible and inaccessible chromatin. Accessible chromatin very often matches with active gene regulation—whether that be gene activation or suppression. You can thus gather much about a cell’s type, state, and regulatory elements from its chromatin accessibility profile.
Download the 10x Genomics Information Guide
The name explained
scATAC-seq is the short form for single-cell assay for transposase-accessible chromatin with high-throughput sequencing. Unpacking that name is a helpful way to understand what the technology entails. So, let’s dissect the name term by term.
- Single-cell assay | Relating to technologies that capture data of individual cells.
- Transposase-accessible chromatin | ATAC-seq pivots around an enzyme called the Tn5 transposase. This enzyme can access open chromatin regions, which are generally sites of active gene regulation. Transposase cuts and inserts 10x Genomics barcodes into the open chromatin in a single experimental step.
- High-throughput sequencing | These adapters can be collected and sequenced using next-generation sequencing. Adapters from individual cells are labeled and amplified to detectable volumes, then pooled and sequenced. This results in sequencing reads. Ultimately, each read corresponds to a specific chromatin accessibility event in a single cell.
Read this blog in which we futher explain how scATAC-seq works
The final product of scATAC-seq is a chromatin accessibility profile. This profile shows the regions of chromatin that were open to the transposase at the time of the experiment. These regions are likely areas of active regulation, so these profiles provide information on individual cells’ gene expression.
Example scATAC-seq data shows aggregate open chromatin peaks of, in this case, 300 single cells. When overlaid to a reference genome, the aggregate profile shows which genes are present in the open chromatin. Adapted from: 10x Genomics.
What scATAC-seq enables
Resolve heterogeneity
Like single-cell RNA sequencing, scATAC-seq reveals variations in chromatin accessibility states in heterogeneous cell populations. With this data, you can identify cell types and subtypes, such as neuron subtypes or cancer cell subpopulations. Even when the differences between cell states are difficult to identify or a cell subtype is present in low cell numbers.
It can do so through unbiased means because it does not require known markers to identify a cell type. Thus, it can provide evidence for a new distinction between cell types or to identify novel cell states. As such, it can give new insights into treatment response or disease.
This discovery of an invasive cancer stem cell population is a case in point. Guilhamon et al. study the chromatin accessibility in glioblastoma cells. They show, using single-cell ATAC-seq, that glioblastomas harbor a heterogeneous self-renewing stem cell population.
They could identify three stem cell states based on their chromatin accessibility profile: Reactive, Constructive, and Invasive. Uniquely essential transcription factors seem to govern each type and the cell states are present in patient glioblastomas in different proportions. Moreover, the stem cell states connect to cancer survival rates in xenograft mouse models. The ratio of stem cell states present in the tumor determined the mouse’s chances of survival.
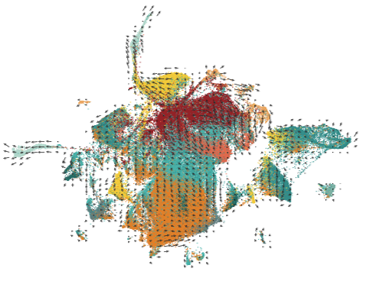
Six cell clusters, identified by scATAC-seq and depicted in UMAP plots, from a glioblastoma sample. Some clusters are enriched in stem cell transcription factor promoters (STEM). Based on their scATAC-seq profile, the individual cells were identified as four known cell types (astrocyte-like, mesenchymal-like, neural progenitor-like, and oligodendrocyte-like). Simplified and adapted from: Guilhalmon et al. (CC-BY 4.0)
Detect dynamic changes in therapy or disease
Researchers can monitor dynamic changes in chromatin accessibility. For example, they can follow the dynamics across different developmental stages to understand how cells alter their gene expression over time. Or they can measure accessibility changes in response to stimuli, for instance in response to therapies that work through gene regulation.
An example is vorinostat. This anti-cancer drug works on histones, chromatin’s protein elements, and alters gene expression by changing chromatin accessibility. Drug developers use ATAC-seq to study the chromatin accessibility changes connected to vorinostat treatment.
Uncover regulatory elements
Identifying the underlying sequences of accessible chromatin gives more insight into gene regulation. With what we know of the targets and DNA binding motifs of regulatory elements, this information helps identify cell-specific regulators. In this way, scATAC-seq offers insights into the enhancers, promoters, and regulatory elements of cell types.
For instance, a team of scATAC-seq pioneers applied the technology to study T cells in the tumor microenvironment after immunotherapy. While some T cells respond well to immunotherapy and start attacking the tumor, others don’t or become exhausted after time. Understanding the different responses brings the opportunity to improve immunotherapy.
Studying basal cell carcinoma, the researchers identified regulators of therapy-responsive T cells from their chromatin profiles. They uncovered a shared regulatory program that governs T cell exhaustion and T helper cell development. That could be useful to advance therapies.
Link epigenetics to gene expression
Researchers can directly correlate chromatin accessibility with gene expression patterns by integrating single-cell ATAC-seq with single-cell RNA-seq data. The results can shed light on the regulatory mechanisms underpinning cellular functions.
An example is this research on digit development. It studies HOX transcription factors, which play various roles in establishing different parts of the body. The researchers found one HOX type, HOX13, to be active in the cells developing into the distal limb.
Single-cell ATAC-seq results showed that chromatin with regions for binding of HOX13 was open in these cells. When they knocked out the Hox13 gene in mice, the chromatin accessibility profile changed into one associated with the proximal limb. The results indicate that HOX13 makes chromatin accessible for other transcription factors, which promotes a proximal-to-distal limb transition.
scATAC-seq at Single Cell Discoveries
We are excited to announce that Single Cell Discoveries has expanded its services to include single-cell ATAC-seq. The version we offer is 10x Single Cell ATAC. With this recent addition, researchers can now access our expertise in single-cell genomics to explore the epigenetic profiles of single cells at scale. Whether you are investigating gene regulation, cell heterogeneity, or the dynamics of chromatin accessibility, scATAC-seq holds potential for your research.
Want to learn more about how to apply scATAC-seq to your specific projects? Don’t hesitate to contact our dedicated team of experts and discuss the possibilities it offers. We support you in supercharging your research with single-cell chromatin accessibility.
Find out more
Find out more information on Single Cell ATAC, our other 10x Genomics services, resources, and much more in our information guide: