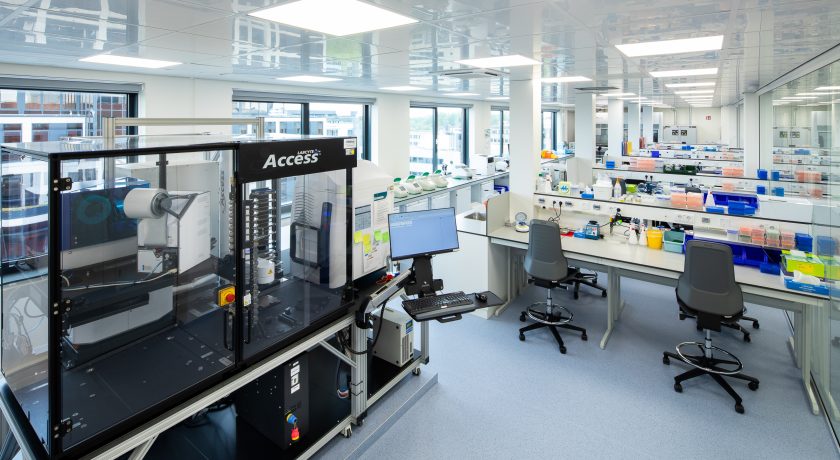
There is one question that should precede almost every other decision in single-cell experiment design: "what is the rarest cell population I really care about?"
Not which platform to use.
Not how many cells your budget allows.
Not what a comparable paper in your field ran.
The rarest population you need to detect — that single number does more to define your experimental design than almost anything else.
Get it right, and cell number, sequencing depth, and platform choice follow naturally.
Get it wrong, and you might either miss the biology you were looking for, or spend significantly more than you needed to find it.
Not sure about cells vs reads?
Frequency determines scale
The relationship between target population frequency and required cell number is straightforward in principle, but routinely underestimated in practice. One of the most common questions we get in our day to day are 1. which technology should I chose? 2. How many cells should I aim for? and 3. What is the best sequencing depth?
If the rare (sub)population you care about represents roughly 10% of your sample, you have a lot of flexibility. Somewhere between 1,000 and 3,000 cells is typically sufficient to detect it with confidence. At this frequency, the choice of platform — plate-based (ie SORT-seq or VASA-seq), droplet/microwell-based (10x Genomics or BD Rhapsody), or combinatorial barcoding (Parse or Scale)— is driven by other factors: sensitivity requirements, sample type, budget. The population frequency alone is not the constraint.
At ~1% frequency, the picture changes. You now need somewhere in the range of 10,000 to 20,000 cells to reliably capture and characterize that population. Plate-based methods start to struggle here — not because of sensitivity, but simply because of throughput. Droplet-based platforms like 10x Genomics and combinatorial barcoding approaches like Parse or Scale are the natural fit: they were built for exactly this scale, and they do it cost-effectively.
At 0.1% or below (for a population you cannot enrich), you are looking at 50,000 to 100,000+ cells, and combinatorial barcoding is often the only realistic path, especially if you are screening many samples. The ability to process very large cell numbers without proportional reagent cost is precisely what these platforms were designed for — and at this frequency, that advantage becomes decisive.
There is, however, an important exception: enrichment. If you can sort or enrich for your rare population before sequencing — via FACS, for example — you fundamentally change the frequency in your input material. A population that is 0.1% in whole tissue might represent 20% or more of your sorted fraction. That shifts you back up the scale, and suddenly plate-based methods are not only viable but potentially advantageous, offering higher sensitivity and full-length transcript coverage that droplet and combinatorial approaches cannot match.
Whether to enrich is therefore a strategic question, not just a logistical one. It is worth asking explicitly before committing to a platform.
Cells and reads are a trade-off, not independent choices
Every single-cell experiment is defined by two numbers: cells captured and reads per cell. Within a fixed sequencing budget, these trade off directly against each other. More cells means fewer reads per cell. This sounds obvious, but it is routinely ignored in experiment planning.
The right balance depends on what you are actually trying to do.
For cell type identification, sequencing depth requirements are lower than most researchers assume. Major cell types are defined by strongly expressed markers that are detectable at modest depth. Sequencing beyond roughly 25,000 reads per cell for a cell typing experiment often delivers diminishing biological returns — budget that would be better spent on additional cells, more samples, or a validation step.
For deep gene expression analysis — co-expression patterns, low-abundance transcripts, subtle differences between similar cell states — depth matters considerably more. Under-sequencing here is a genuine risk. The signal simply isn't in the data if you didn't sequence deeply enough to capture it.
The practical tool for navigating this is the sequencing saturation curve. Saturation tells you the point at which additional reads stop yielding new unique transcripts. If you are near saturation at your planned depth, sequencing more will not give you more biology. If you are far from it, you may be leaving signal on the table. A pilot experiment — even a small one — gives you that curve for your specific sample type, and it is one of the most useful things you can have before committing to full-scale sequencing.
When the honest answer is: I don't know yet
The rarest population question only works if you can actually answer it. Often, you cannot — at least not with confidence. You may have a hypothesis, but not a reliable frequency estimate. You may be working with a sample type you haven't profiled before. You may not know how many distinct cell populations your tissue contains, let alone what fraction each represents.
This is not a planning failure. It is simply reality for a large proportion of real discovery experiments. And it is exactly the situation a pilot is designed to resolve.
A well-designed pilot tells you the cell type composition of your sample, gives you frequency estimates for the populations you care about, generates a saturation curve that guides your depth decisions, and confirms that your sample handling and preparation workflow is working as intended. If the pilot surfaces a problem — poor viability, an unexpected cell type bias, a protocol incompatibility — you find out early, when it is still cheap to fix.
The pilot is not a delay. It is the thing that prevents you from running a large, expensive experiment designed around wrong assumptions. In our experience, the cost of a good pilot is almost always returned many times over in the quality of the full experiment that follows.
Pilot early. Pilot often. If a pilot cannot be performed, or if you simply need a quick proxy for a budget calculation, ask yourself: how complex is my tissue of choice. If you expect only 2 or 3 abundant cell types, 1000 cells might already be enough. If you are studying a complex tissue with lots of likely rare subpopulations (for example brain) a higher number is a good starting point.
Putting it together
Before finalizing your experimental design, it is worth working through these questions explicitly:
- What is the rarest population you really care about, and what frequency do you expect it to be present at?
- Can you enrich for it? If yes, does that open up platform options you hadn't considered?
- Are you cell typing, or doing deep expression analysis? That answer sets your sequencing depth target.
- Do you have a saturation curve for your sample type? If not, is a pilot the right next step?
- Are cells and reads balanced for your actual question, or are you defaulting to what you have seen in the literature?
More cells is sometimes exactly the right answer. Large-scale tissue atlases, high-throughput drug screening, and rare population discovery in complex tissues genuinely benefit from the throughput that combinatorial barcoding platforms offer. But running more cells than your biology requires — because it feels safer, or because a published study used that number — is one of the most common ways to spend money without gaining anything in return.
The best experiments are not the biggest ones. They are the ones where cell number, sequencing depth, and platform choice were each decided for a reason.
At Single Cell Discoveries, experimental design is part of every project we take on. If you are planning a single-cell experiment and want to think through the right approach for your question, get in touch!
Download our services brochure