Single-cell sequencing can be expensive, especially compared to conventional bulk RNA sequencing methods. But there is a good reason why this technology is expensive, and you can mitigate these costs if you carefully consider the design of your experiment.
This post looks at the factors affecting the cost of your single-cell sequencing project, and how adapting the design of your experiment can keep costs down.
The costs of the reagents
For a single-cell sequencing experiment, we process single cells in individual reactions, each of which requires its own set of reagents and this drastically increases the total amount of reagents needed.
In contrast, when we analyze pooled cells in a single tube for a bulk RNA-seq experiment, we use much less reagent per cell. Generally speaking, the reagent costs are 10-20 times higher for a standard single-cell RNA sequencing experiment than that of a bulk RNA experiment.
For single-cell sequencing, reagents costs are 10 to 20 times higher than for traditional bulk RNA sequencing.
The cost of sequencing
The cost of sequencing itself is often overlooked when considering the price of a single-cell sequencing project.
Just as with the reagents needed for library preparation, you will need many times more sequencing reads for a standard single-cell sequencing project than for a standard bulk RNA sequencing project.
A typical bulk RNA sequencing analysis requires up to 20 million sequencing reads per sample. In contrast, the typical number of reads per cell needed for a single-cell sequencing project ranges from 50,000 to 150,000*.
So, for example, a relatively simple single-cell sequencing project on the 10x Genomics platform, where you target 3,000 cells with a sequencing depth of 50,000 sequencing reads per cell, will need to generate 3,000 x 50,000 = 150 million reads. That's 7.5 to 15 times more than for a bulk RNA analysis!
Sequencing costs for single-cell sequencing are approximately 7 to 15 times higher than for traditional bulk analyses.
But could you use fewer reads per cell?
Sure, you could. But to gain a thorough understanding of the transcriptome at the level of the individual cell, you need adequate sequencing depth to get a statistically significant dataset.
This is one of the most important trade-offs to consider if you want to keep the sequencing costs stable. If you double the number of cells in your analysis, you will have to halve the number of reads per cell.
The biological question you want to answer will determine which factor is most important.
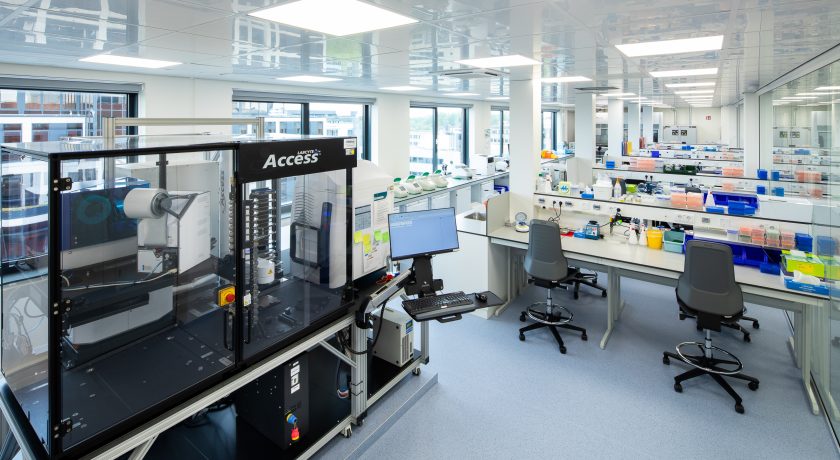
High-throughput single-cell sequencing is performed on the 10x Chromium X
The technology
Single-cell sequencing is high-tech. It’s a relatively new technology that only recently became available to a larger market thanks to a reduction in costs, as well as innovations in scaling up and standardization.
But, whatever platform or technology you choose, it is more sophisticated than conventional molecular biology technologies such as bulk RNA sequencing.
In order to be able to offer single-cell sequencing in a commercially viable way, we needed to scale down the volume of the reagents used. This could only be done with sophisticated and expensive liquid handling robots.
The same goes for the microfluidics devices that use beads. These beads are microscopic and each of them needs to be coated with a molecular barcode. For one sample, up to 3 million unique barcodes are required. Furthermore, these beads have to run through a microfluidics device smoothly and consistently.
All this requires active research and development, which all adds to the cost of single-cell sequencing.
The setup of your experiment
The most important factor in the cost of single-cell experiments is your biological question. The ultimate price of a project depends on 3 important factors: the number of samples, the number of target cells per sample, and the number of reads per cell.
Read our eBook on Single Cell Sequencing in Drug Development
Get inspired by successful drug developers employing single-cell sequencing in their research; a review of 10 years single-cell sequencing.
Number of samples
The math is simple: the more samples, the more your project will cost. This is true of all projects, but especially so for single-cell sequencing projects.
And this is why it is so important to take a critical look at the set-up of your experiment: Do you really need to include every single patient from your study? And do you really believe that those 7-time points will all be different?
If you take a second look at the setup of your experiment, you may realize that you don't need to analyze as many samples as you first thought. It may even be possible for you to achieve your aims if you combine single-cell experiments with much cheaper bulk RNA experiments.
Or you could opt for a step-wise approach, start small, and scale up later if the data tells you to.
Number of target cells per sample
Again, the math is simple: more cells means a more expensive project - unless you trade this off against fewer reads per cell. With the plate-based SORT-seq system, this is because for every additional 384 cells, you'll need to add another plate to your experiment.
The cost of each plate consists of a fixed part (reagent and handling costs) and a variable part (sequencing costs).
With the 10x Genomics system, where more cells are typically processed per sample, the sequencing costs can be a significant proportion of the total price.
So, if you process tens of thousands of cells for one experiment, the price will rise significantly. Again, think about your project and the biological question or hypothesis you want to answer.
If, for example, you want to study 4 different cell types in a tissue sample, and you expect them to be present in more or less equal ratios, then you probably don't need many thousands of cells per sample.
However, if you are interested in a very rare cell type that you expect to occur in only about 1% of your sample, then you will need to target thousands of cells to properly analyze the cell type you're interested in.
Number of sequencing reads per cell
Once again, the math is simple here: more sequencing reads increases the costs of your experiment. As we said earlier, the total number of sequencing reads required per sample for single-cell sequencing can be quite high.
So again you need to be pragmatic about your experiment, biological question, and hypothesis.
If you want to call the cell types in a sample, there is no need to sequence very deeply (e.g. >50,000 reads per cell). But if you are interested in a very specific, lowly expressed gene from a certain pathway, then you might want to increase the sequencing depth (e.g. 100,000 or 150,000 reads per cell).
We can help you with these decisions. Our staff have extensive experience with setting up single-cell sequencing projects and can help you find the best set-up for your project.
Schedule a call with one of our specialists today to discuss your project and to see how we can help you keep your costs down.
*Minimum recommended sequencing depth is 20,000 reads per cell.
Ready to proceed? Use the form below to download our information guide.