T cells don’t just patrol the immune system; they remember, recognize, and respond. At the center of this ability is the T cell receptor (TCR), a molecular antenna tuned to detect antigens (depicted on the right). For years, deciphering the behavior of individual T cells was like trying to follow a conversation in a noisy room: possible in fragments, but far from clear.
Advances in TCR-coupled single-cell RNA sequencing have changed that. This approach now allows us to capture each T cell in high resolution, connect its transcriptional profile to its unique TCR sequence, and map it within the broader immune landscape.
In this post, we will guide you through a comprehensive analysis journey, from UMAP clustering to clonotype tracking and epitope annotation, revealing how T cells behave, interact, and respond to their environment.

Jump to a section in this blog:
- Map The T Cell Landscape
- Identify Cell States with Marker Genes
- Measuring Expansion: Clonotype Frequency Across Transcriptional T Cell Clusters
- Visualize TCR Similarity Clusters
- Projecting Annotated Epitopes Onto the UMAP
- Putting It All Together: Epitope Specificity Within Clonal Groups
- Wrap-Up: Why This Analysis Matters
- Interested in What Your T Cells Are Really Seeing?
Map the T Cell Landscape
TCR-coupled single-cell RNA sequencing is part of a broader field known as immune cell receptor profiling, which involves the study of adaptive immune receptors (TCRs and B cell receptors, or BCRs) at scale. T cell repertoire profiling enables us to measure the diversity, clonal expansion, and antigen specificity of TCRs across timepoints, tissues, or disease states.
At Single Cell Discoveries, we offer an end-to-end TCR single-cell sequencing solution (utilizing 10x Genomics 5’ TCR Immune Profiling), starting from sample processing and library preparation, through immune repertoire profiling and advanced bioinformatics, to deep insights into clonal expansion, T cell phenotypes, and receptor diversity, all guided by expert data consulting to help you interpret and translate findings into discovery. Powered by ImmuneWatch DETECT, we now offer TCR-epitope annotation capabilities, unlocking the ability to infer what your T cells are actually recognizing.
Get the 10x Genomics
information guide
You can find a full explanation of 10x Genomics, its variations, and our approach in our information guide.
ImmuneWatch DETECT is the leading software solution for T cell receptor specificity annotations. It utilizes an extensive, carefully curated database of TCR-epitope pairs and a public benchmark-winning algorithm to provide the most accurate epitope specificity annotations. Knowing the importance of your data, ImmuneWatch DETECT was built with data privacy at its core. All processing is done locally, ensuring your valuable information never leaves your machine.

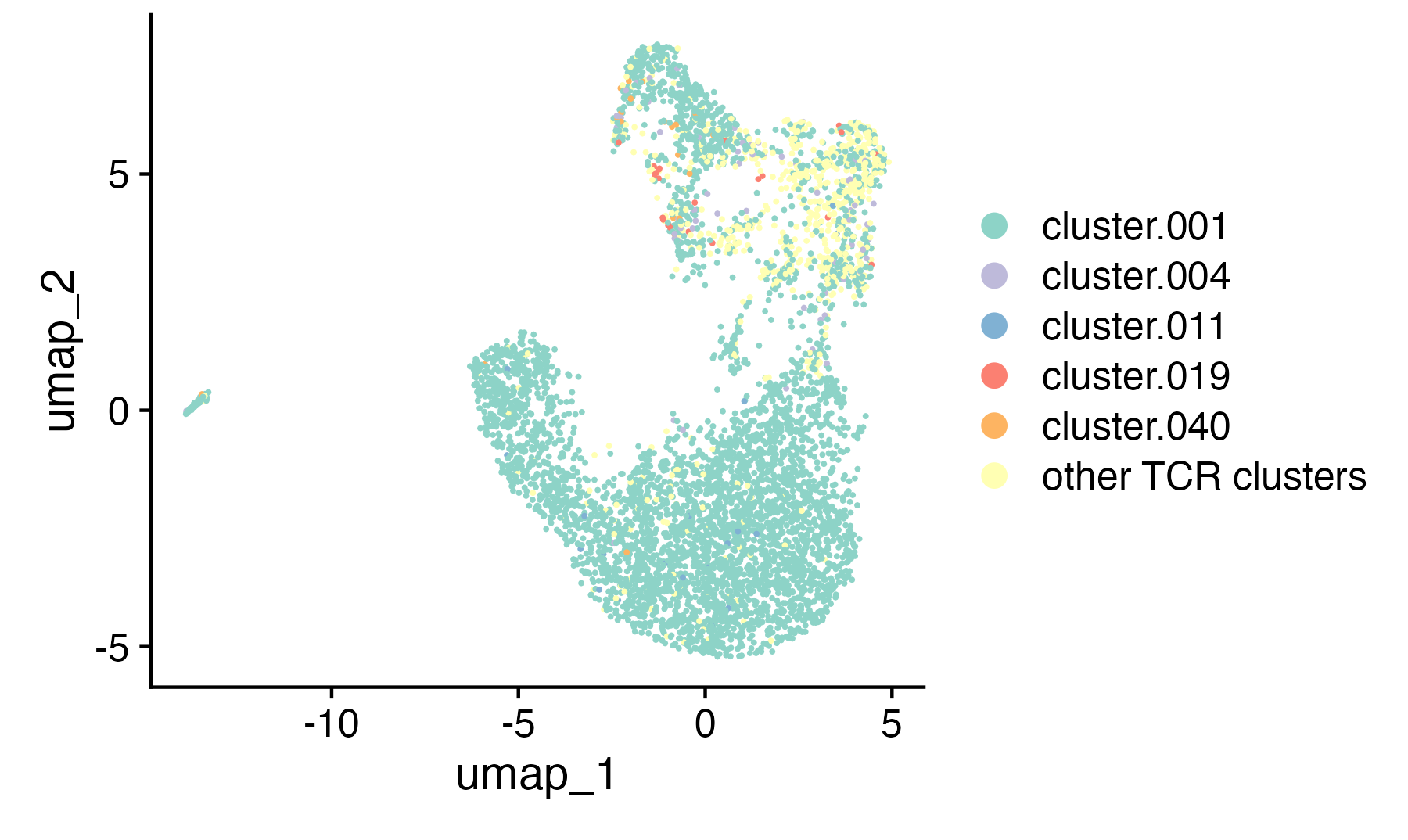
UMAP projection of single-cell T cells colored by transcriptional cluster.
We start with transcriptomic data and project our T cells into low-dimensional space using UMAP. Each point is a cell; clusters reflect distinct transcriptional identities.
Here, you can already see structure emerge: some clusters are tightly packed, others more diffuse, depending on the transcriptional similarity of neighboring cells.
Identify Cell States with Marker Genes
To give biological meaning to each cluster, we overlay key marker genes.
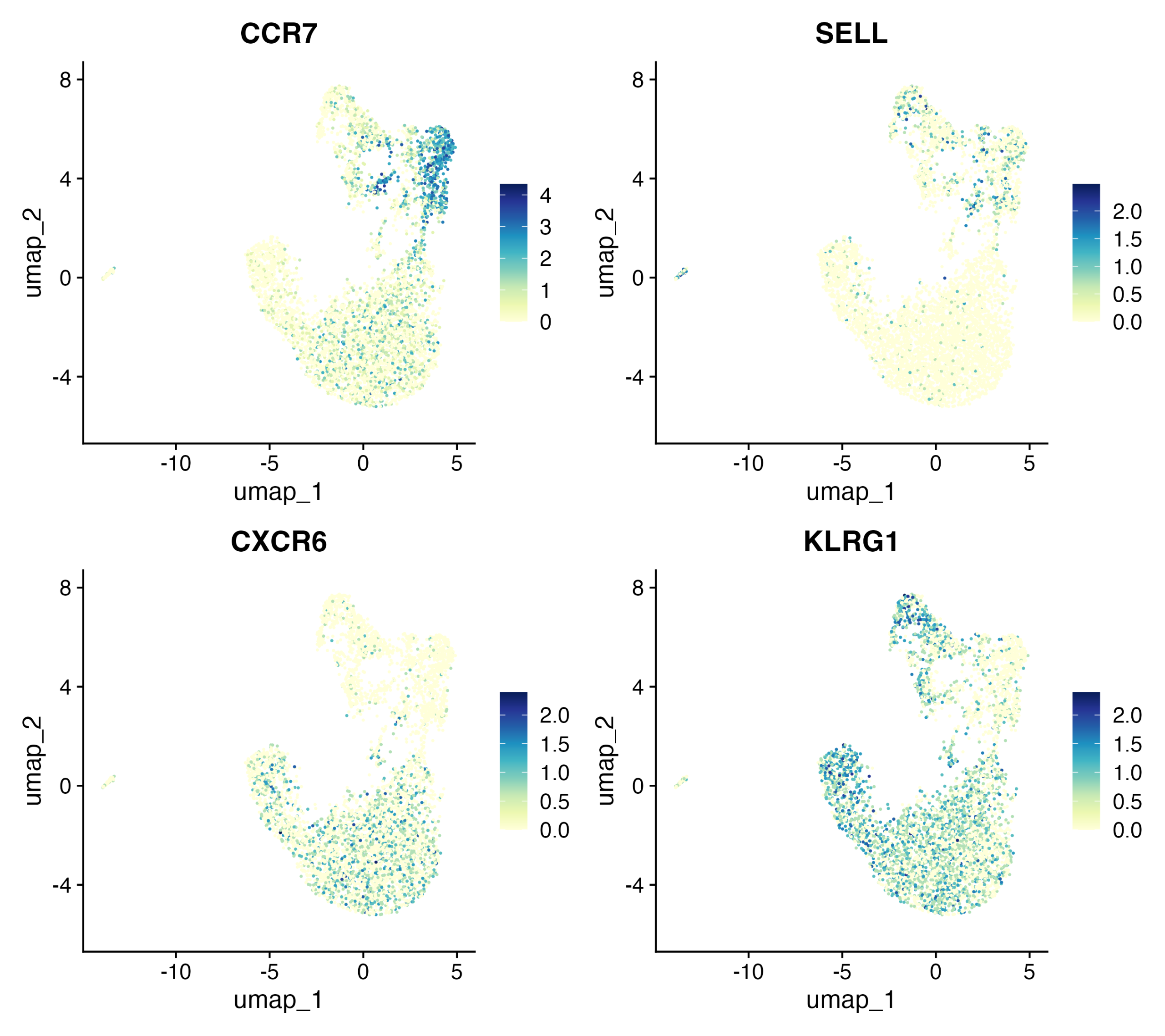
Expression of canonical markers such as CCR7, SELL, CXCR6, and KLRG1 across the UMAP.
These markers help annotate clusters as:
- Naïve (CCR7)
- Naïve/Central Memory (SELL)
- Tissue-homing/Effector (CXCR6)
- Effector/terminal (KLRG1)
Measuring Expansion: Clonotype Frequency Across Transcriptional T Cell Clusters
A TCR clonotype refers to a group of T cells that share the same T cell receptor sequence, typically defined by identical or highly similar CDR3 (Complementarity-Determining Region 3) regions. These cells result from a convergent response and often recognize the same antigen. Tracking clonotypes allows us to study clonal expansion, immune diversity, and antigen-specific responses during infection, cancer, or autoimmunity. We can quantify how expanded each clone is and where it's located.
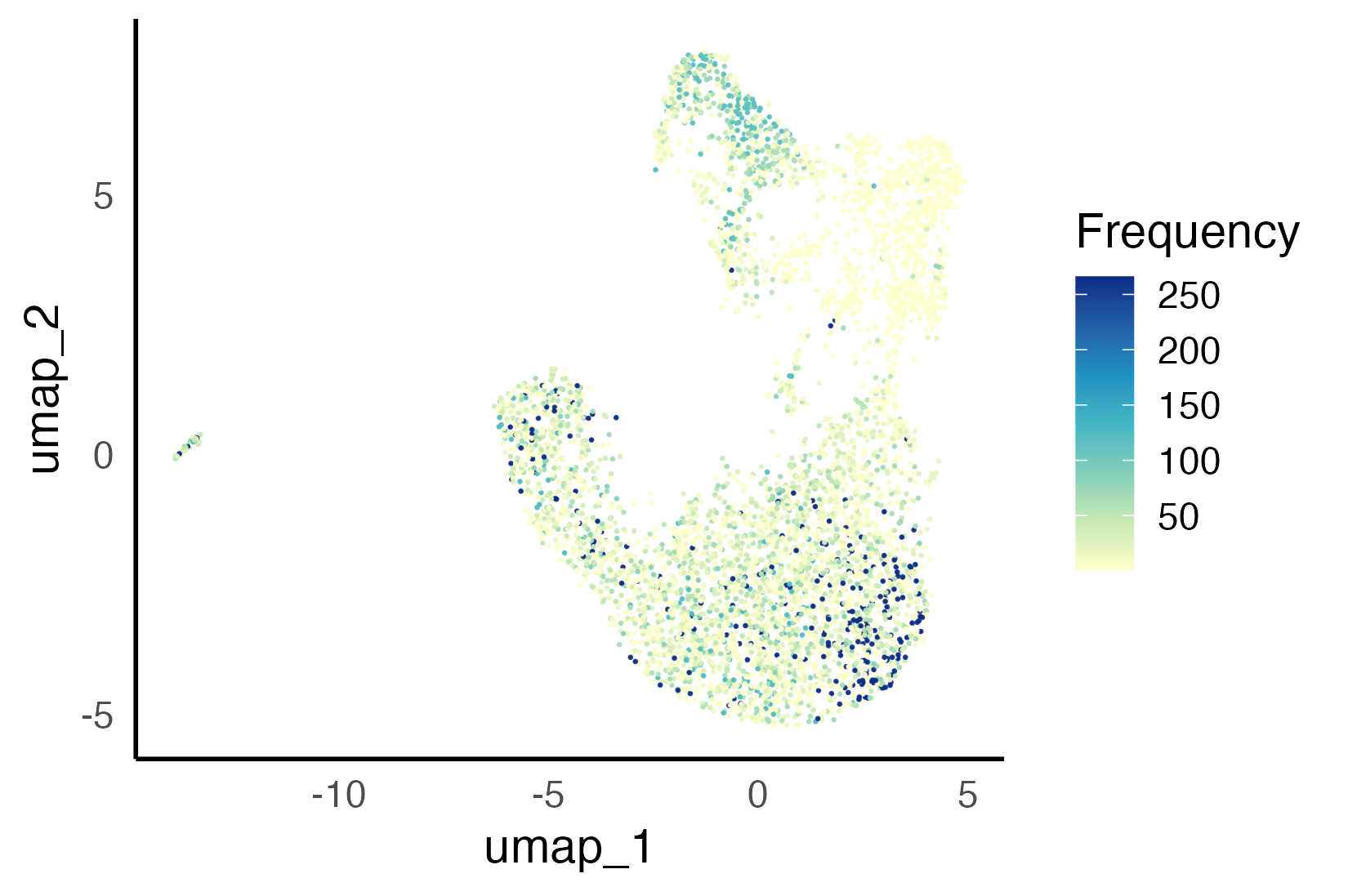

UMAP with clonotype sizes visualized (left), and bar plot of clonotype frequency per transcriptional (i.e. clustered on gene expression) T cell cluster (right).
Study an example Exploratory Data Report
Explore our Exploratory Data Report for a real dataset, demonstrating how we connect count tables to t-SNE and UMAP visualizations, and take the first steps toward cell type annotation.
Focus on the Top 10 Clonotypes
To really get a feel for the immune response, we highlight the top 10 most expanded clonotypes.

UMAP showing the distribution of the top 10 clonotypes across cell clusters.
You will notice that some clones dominate one cluster, while others are scattered across several. That can reflect clonal plasticity, or ongoing differentiation.
Visualize TCR Similarity Clusters
We can also group cells with identical or highly similar TCR sequences into TCR similarity clusters.

Pie chart of TCR similarity clusters (left) and their spatial distribution on the UMAP (right).
You might see dominant TCR similarity clusters spreading across transcriptional states — or grouping in a single phenotype. A clonal T cell in an effector cluster? That could be tumor-reactive. A diverse polyclonal naïve cluster? Maybe just immune surveillance.
What Are They Responding To?
Using ImmuneWatch DETECT, we can annotate which epitopes the clonotypes are likely to recognize. Internally, the epitope prediction tool will compare the clonotype sequences with its own database of TCR-epitope pairs and will output a score on how likely this specific clonotype will bind to a certain epitope. It will also predict the MHC allele to which the TCR might be specific and provide a binding score for the predicted MHC allele. Additionally, the tool will provide the antigen and species name to which this epitope belongs, along with a link to the published source on which the annotation is based.

Example output of ImmuneWatch DETECT, of which the columns highlighted in blue are added to the TCR repertoire file by the annotation tool. Input TCR sequences are not used for training ImmuneWatch DETECT.

Top 50 annotated T cell epitopes by abundance (based on TCR binding likelihood via ImmuneWatch DETECT).
This plot reveals whether viral, tumor, or autoimmune-related epitopes dominate a sample. One might find:
- Viral epitopes (e.g., SARS-CoV-2 spike)
- Tumor antigens (e.g. MART-1, KRAS, NY-ESO-1)
- Self-peptides (for autoimmune studies)
Projecting Annotated Epitopes Onto the UMAP
Now, we can observe how those annotated epitope specificities map back onto the cells.
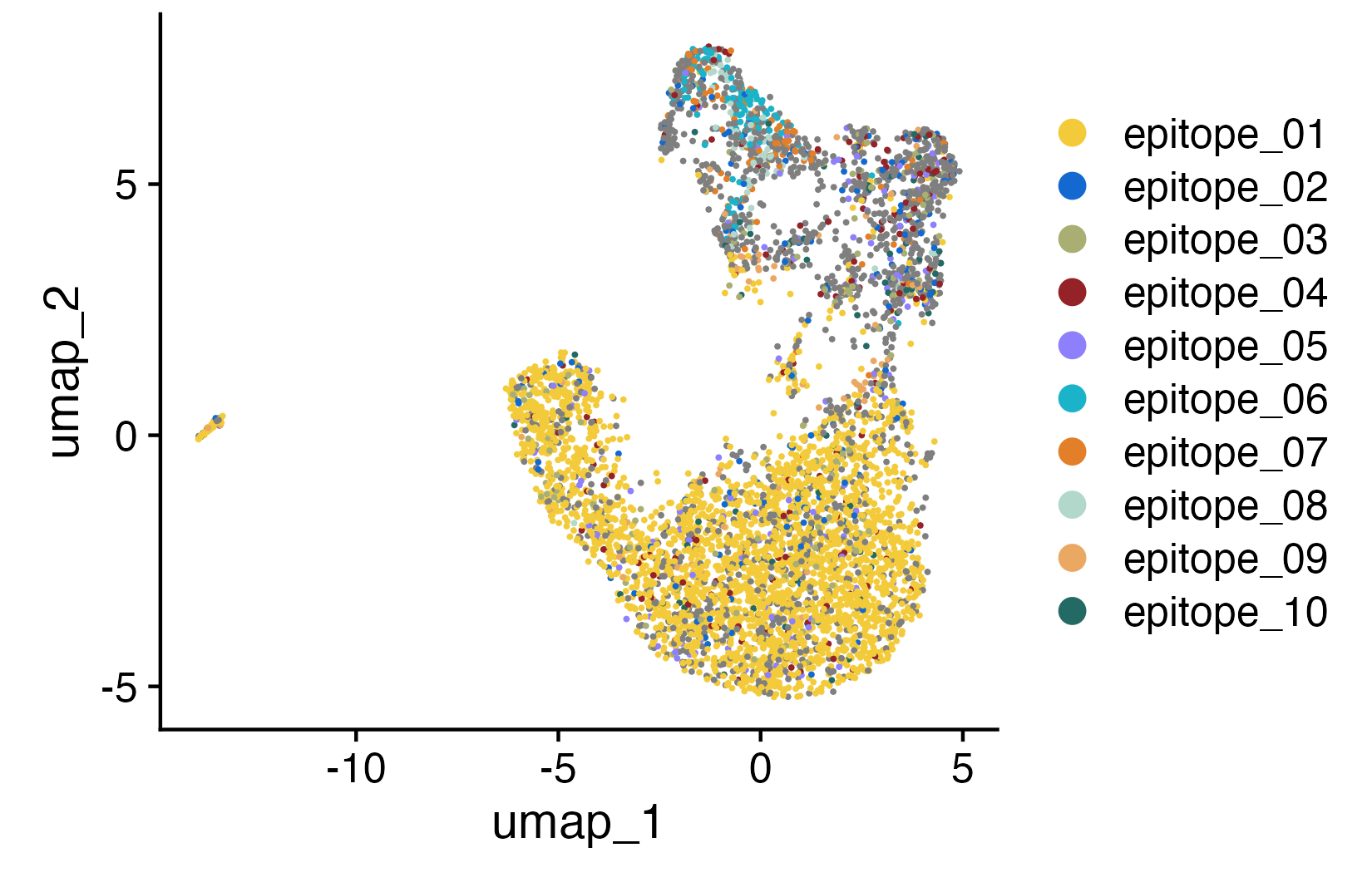
UMAP colored by annotated epitope binding per cell (left) and TCR clonal clusters colored by annotated epitope binding (right).
Patterns may emerge: entire clusters reacting to the same epitope, or a single epitope recognized across diverse cell types. Either way, you're linking T cell receptor, function, and antigen in one shot.
Putting It All Together: Epitope Specificity Within Clonal Groups
Finally, we explore which epitopes are annotated for each clonotype within TCR similarity clusters.


(Left) A clonal network where each node represents a unique TCR, and the node size indicates the number of cells sharing this TCR. Nodes touching or connecting (represented by gray lines) represent TCR similarity clusters. Nodes are colored by annotated epitope specificity within TCR similarity clusters. (Right) Top 100 clonotypes matched to annotated epitopes with connecting lines colored by TCR similarity cluster.
This demonstrates epitope convergence, where different TCRs converge on the same specificity, often through shared motifs. You might spot:
- Multiple clones targeting the same viral antigen
- Shared epitope specificity in different patients
- Broad vs. narrow epitope targeting patterns
Wrap-Up: Why This Analysis Matters
Whether you're studying COVID-19 responses, tumor-infiltrating lymphocytes, or designing TCR-based therapies, single-cell TCR analysis lets you:
- Decode what T cells are doing
- Understand how they are changing
- Predict what they’re responding to
This integrated approach offers an unprecedented window into the immune system — one that’s high-resolution, interpretable, and incredibly powerful.
Interested in What Your T Cells Are Really Seeing?
At Single Cell Discoveries, we offer an end-to-end TCR single-cell sequencing solution: from immune repertoire profiling (using 10x Genomics 5’ TCR Immune Profiling) to advanced bioinformatics and epitope prediction, giving you deep insights into clonal expansion, T cell phenotypes, and antigen specificity. Whether you're studying cancer, infection, or autoimmunity, we deliver actionable, publication-ready data that connects immune receptor identity with function and predicted targets.
Want to take a deeper dive into your data? We offer extensive data consulting services, covering comprehensive QC, normalization, and clustering, dimensionality reduction, differential expression, pathway analysis, and immune repertoire profiling, all tailored to your dataset and research goals through expert consultation. Our offering now also includes ImmuneWatch DETECT, which provides epitope annotations of your TCRs, giving you the best possible picture of what your cells are doing.